Beyond the Magic Bullet: Why the Future of ADCs is Written in Spatial Biology
Antibody-drug conjugates (ADCs) represent one of oncology’s most promising frontiers. Despite decades of research, fewer than 15 FDA-approved ADCs exist, and clinical trial failure rates remain above 90% from phase 1 through phase 3. The promise of targeted therapeutics has long captured the imagination of oncology, but the reality is more complex: tumors aren't just clusters of malignant cells wanting to be eliminated. They are intricate, dynamic ecosystems with defensive architectures that conventional ADC design has struggled to navigate.
At Owkin, we aren't just building better molecules; we are building navigation systems that understand the 'where' and 'why' of the tumor before we ever design the 'how'.
The ADC Evolution: From Random Chemistry to Precision Engineering
A Brief History of ADC Generations
The evolution of ADCs has progressed through distinct technological waves:
- First-generation (1990s-2000s): characterized by unstable linkers and stochastic (random) payload attachment, leading to unpredictable drug release and systemic toxicity.
- Second generation (2000s-2010s): Introduction of cleavable linkers and more potent payloads, but still limited by off-target effects.
- Third generation (2010s-present): site-specific conjugation technologies enabling more consistent drug-antibody ratios and improved therapeutic windows.
The Persistent Challenge
Despite these advances, traditional ADCs often hit a clinical "glass ceiling" caused by two interlocking failures:
- The Narrow Therapeutic Window: Because traditional targets are often shared with healthy tissues, we are forced to limit the dose to avoid systemic toxicity.
- Adaptive Resistance: This "under-dosing" allows the most resilient cancer cells to survive and evolve. Whether through target down-regulation or spatial exclusion - where the tumor microenvironment physically shields cancer cells from the drug - resistance mechanisms quickly render standard ADCs ineffective.
At Owkin, we believe the solution to both is Biological Context. By using spatial data to identify precision target strategies—whether bispecific co-localization, heterogeneity-aware targeting, or context-specific delivery—we widen the therapeutic window. And by mapping how tumors adapt under previous treatment pressure, we anticipate resistance, designing ADCs that stay ahead of the tumor's ability to evolve.
The Cost of Failure
Late-stage ADC failures represent more than setbacks in drug development, they signal fundamental gaps in our understanding of tumor biology and therapeutic delivery. Each Phase II or III failure due to toxicity or lack of efficacy forces difficult decisions about portfolio prioritization, not to mention the impact on patients who continue to await better treatments.
At Owkin, we believe the next generation of ADCs isn't defined by a stronger toxin, but by superior biological context. While traditional ADC design relies on bulk surface-level expression, we use high-resolution spatial and multi-omic data to map the "geography" of the tumor, transforming ADC development from a game of trial-and-error into a discipline of predictive engineering.
The Owkin Difference: Spatial Biology-First ADC design
The three-pillar approach described below is operationalized through Owkin’s AI Scientist, K Pro. K Pro connects researchers directly to the MOSAIC dataset, enabling natural language queries across the entire ADC design workflow: from identifying co-localized receptor pairs (Pillar 2) to mapping tumor microenvironment enzymatic signatures (Pillar 3) to predicting how target expression evolves under treatment pressure (Pillar 1). Through these capabilities K Pro transforms spatial biology into an integrated drug discovery engine accessible to all our researchers.
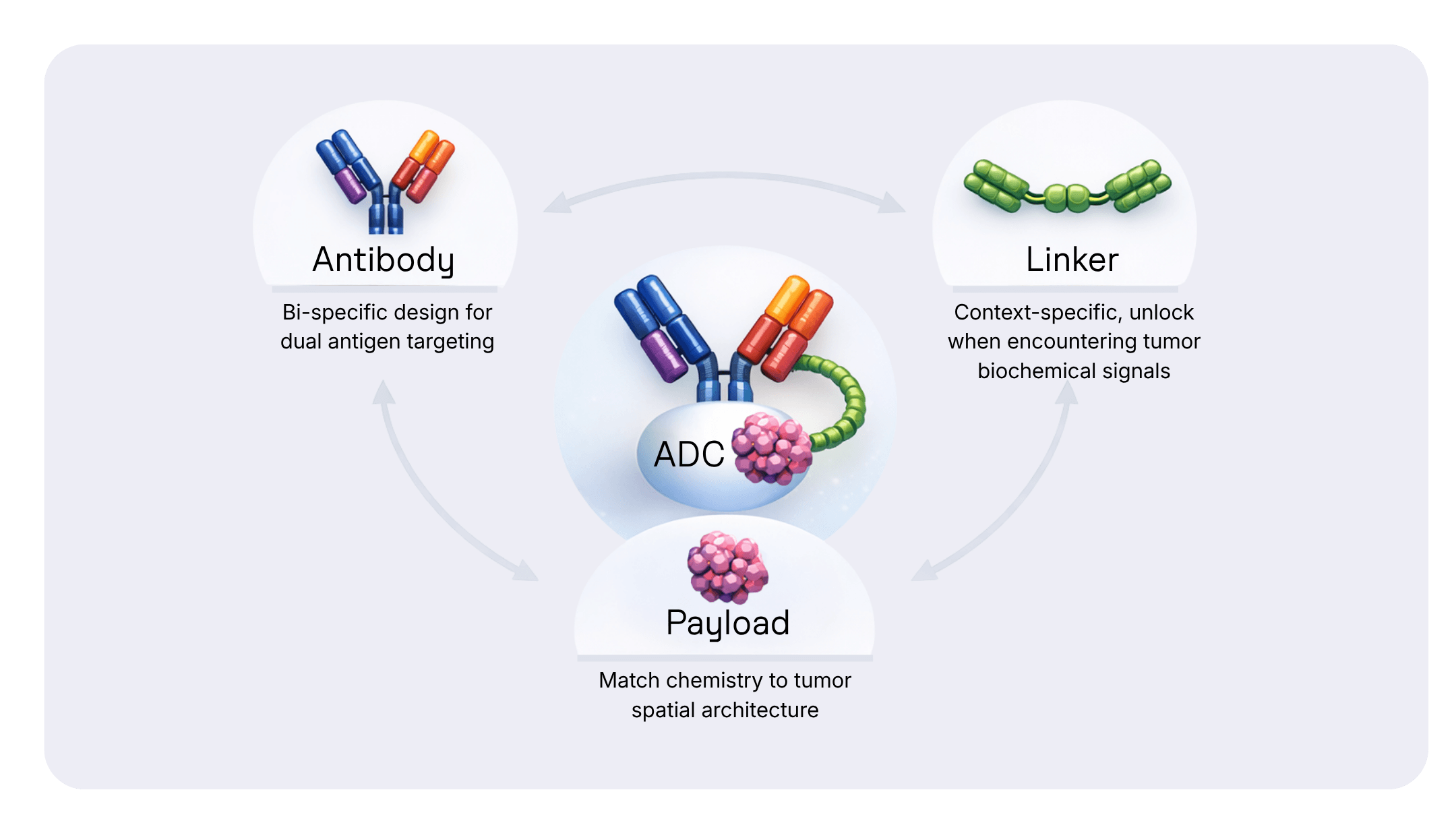
Pillar 1: Multi-omic 360° Intelligence
Traditional ADC design asks: "Is the target present?" We ask: "Where is it? At what levels? What is it “talking to”? And how will the surrounding environment respond?"
This requires a high-fidelity, multidimensional map of the tumor’s geography before designing a new ADC. Through the MOSAIC (Multi-Omics Spatial Atlas in Cancer) initiative, we have profiled 2000 patients across 7 cancer types with spatial and multi-omics data2. From this foundation, our AI-powered discovery platforms integrate six complementary data layers:
- Digital Pathology / H&E: Analyzing tissue morphology and architecture
- Bulk RNASeq: Quantifying overall gene expression patterns and tumor-level signatures
- Whole Exome Sequencing (WES): Identifying genetic drivers and mutations
- Single-Cell RNA-seq: Decoding individual cell states, types, and transition dynamics
- Longitudinal Clinical Data: Understanding real-world patient outcomes, treatment response, and resistance patterns
- Spatial Transcriptomics: Mapping gene expression in its spatial tissue context, revealing how cells communicate with their neighbors and how targets are distributed within the tumor microenvironment
By layering these datasets, our AI platforms can identify not just which targets are present, but how they are distributed spatially and across different cell-types (critical for predicting payload penetration), how they interact or co-express with other factors (essential for bispecific pairing and linker design), and how they evolve across disease states (key to anticipating resistance).
Specifically, we have developed a series of methodologies that enable us to:
- Spatially map the proximity of targets to immune infiltrates, allowing us to predict which ADCs will most effectively prime the tumor microenvironment for the combination of ADCs with Immuno-oncology drug success3.
- Integrating histology with spatial transcriptomics allows us to pinpoint specific zones of tumor infiltration. We use this to identify targets that are expressed in the most aggressive "prognostic hotspots," ensuring our ADCs strike the tumor where it is most vulnerable4.
- Predict patient-level outcomes from spatial patterns, using machine learning to connect tissue architecture with clinical endpoints, enabling biomarker-driven patient stratification5.
- Through spatially integrated multi-omics, analyze how cancer cell populations evolve under the selective pressure of prior lines of treatment. This allows us to match payloads to specific cell states and predict which tumor regions are likely to develop resistance, ensuring our ADC payloads are selected to bypass these cellular defenses1.
- Deconvolve spatial cellular niches to identify receptor pairs that are truly co-expressed and co-localized on the same malignant cells (COMPOTES method3). This avoids the false positives of bulk and single-cell data, providing a high-confidence roadmap for bispecific dual-lock strategies that maximize tumor selectivity.
Pillar 2: Precision Target Selection
Traditional single-target ADCs face a fundamental limitation: any protein expressed on tumor cells is almost always present at some level in healthy tissue. This creates a notorious dilemma: traditional ADCs are often hindered by a binary limitation, where they either suffer from 'on-target, off-tumor' toxicity that narrows the therapeutic window, or they require dose reductions that ultimately compromise efficacy.
The bispecific antibody approach, already validated in therapeutics like Blinatumomab (in Acute Lymphoblastic Leukemia) and Amivantamab (in Non-Small-Cell-Lung-Cancer), offers a solution: require two molecular "keys" to unlock the drug's activity. But identifying the right key pair requires answering critical spatial questions that traditional bulk-RNA-seq or immunohistochemistry (IHC) cannot address easily:
- Are both targets expressed in the same cell, or just in the same tissue?
- Are they co-expressed in tumor-adjacent immune cells or stroma, creating false positives?
- How consistent is this co-localization across different patients and disease stages?
- When targets appear on different cancer cell populations, how can we ensure comprehensive tumor coverage?
This is where spatial biology becomes essential.
Bispecific “Dual-Lock” Targeting
By leveraging our spatial transcriptomics data, we identify pairs of receptors that are exclusively co-localized in the same malignant cell in tumor tissue but remain separate in healthy organs. This dual-lock mechanism ensures the ADC only binds and internalizes when both keys are present, dramatically widening the therapeutic window.
Our multi-omic platform enables us to move beyond simple co-expression to true spatial co-localization, and answer questions like “Which antigen pair provides the highest tumor-specific density?” and “How much of the total tumor volume is 'covered?" by our dual-target strategy?”. This allows us to confirm that both targets are present on the same malignant cell, quantify what percentage of tumor cells co-express both targets at therapeutically relevant levels, and map healthy tissue exclusivity to minimize toxicity risk.
A key advantage of dual-lock targeting is resilience against adaptive resistance. Cancer cells under selective pressure often downregulate targets. By requiring two targets, we force the tumor into a molecular corner where escape becomes far more difficult. Through the MOSAIC dataset, we can track how target expression patterns evolve across disease stages and treatment exposure, allowing us to prioritize target pairs that remain co-expressed across clonal evolution, and show minimal compensatory downregulation in resistant disease.
Covering Tumor Heterogeneity: Harnessing the Bystander Effect
Even with perfect dual-lock specificity, many tumors present a different challenge: intra-tumoral heterogeneity, where different cancer cell clones express different target profiles. A purely co-localized dual-lock strategy might miss significant portions of the tumor mass.
By mapping the spatial distribution of candidate targets across the tumor microenvironment, we identify scenarios where Target A and Target B mark spatially adjacent but distinct cancer cell populations. In these cases, we design bi-specific ADCs that leverage the bystander effect to achieve comprehensive tumor coverage.
By integrating spatial transcriptomics with gene expression profiling, we can:
- Predict payload sensitivity for specific tumor cell clusters before selecting the cytotoxic agent
- Match payload chemistry to the tumor’s spatial architecture, i.e. selecting membrane-permeable agents for targets that are expressed heterogeneously on tumor cells
For every heterogeneous tumor scenario, our spatial platform answers the critical question: is the spatial distribution of targets conducive to the selected payload's diffusion radius?
This turns the tumor’s complexity against itself, ensuring a "total kill" even in the most diverse and heterogeneous malignancies. The bystander effect has been clinically validated in HER2-targeting ADCs treating HER2-low breast cancer, demonstrating that strategic payload selection enables treatment of heterogeneous tumors.
In-Silico Toxicity Profiling
Before a single molecule is synthesized, we conduct in-silico toxicity profiling across our vast atlas of healthy and diseased tissues.
The process looks like this:
- Map target expression across 100 tissue types from our organ atlas
- Simulate the ADC’s journey through diverse cellular environments
- Predict potential "off-target" interactions using AI models
This allows us to refine or discard candidates in days rather than months of costly lab work. Our approach front-loads safety assessment, dramatically reducing time and cost in the discovery phase.
Pillar 3: Context-Aware Engineering
The “Intelligent” Linker: Engineering for the Microenvironment
A tumor is not an isolated mass but a guarded fortress with immune cells, blood vessels, and signaling molecules. Traditional ADC linkers are biochemically "blind"; they release payloads based on general cellular conditions (e.g. pH, protease activity) that might also exist in healthy tissue.
By mapping the TME’s enzymatic signatures with high-resolution spatial data, we engineer context-specific linkers that only "unlock" when encountering tumor-specific biochemical signals1.
Designing for Combination: ADC + IO and Beyond
The future of oncology is not a single Magic Bullet, but the strategic combination of modalities. We are seeing a paradigm shift where ADC + Immuno-Oncology (IO) combinations, such as the recent standard of care successes in urothelial cancer, are redefining patient outcomes.
At Owkin, we use spatial biology to design these combinations with predictive intent:
- Spatial analysis of the TME allows us to identify the optimal combinations of TME-targeting treatments, such as IO and stromal-targeting drugs, with ADC.
- We can combine the analyses of spatial ADC features presented above with spatial features relevant for TME-drugs, and predict both additivity and synergy in specific patients subgroups.
Beyond IO and TME, our platform analyzes the spatial and multi-omic landscape to identify synergies with DNA Damage Response (DDR) inhibitors, anti-angiogenic agents, and other targeted therapies, matching the drug's mechanism to the tumor's specific environmental weaknesses.
Towards a 4th Generation of ADCs
The history of ADC development has been defined by the pursuit of the "magic bullet." At Owkin, we recognize that precision is only as valuable as the biological intelligence guiding it. By anchoring our design process in high-resolution, large-scale spatial biology and multi-omic integration, we are moving past the era of trial-and-error. This depth and scale allow us to apply our insights to any target and any drug combination of interest with unprecedented flexibility. Powered by K Pro's ability to analyze multi-omic patient data, our approach doesn't just ask if a target is present; it asks:
- Where is it located within the tissue?
- Who co-localized in the tumor core?
- How will the surrounding biology respond to therapeutic intervention?
- What resistance mechanisms might emerge, and can we anticipate them?
This systems-level strategy enables us to:
- De-risk candidates through in-silico toxicity screening
- In-silico screening of ADC combinations with IO and other drugs to identify synergistic potential and optimal treatment sequencing before entering the clinic.
- Refine targeting with bispecific precision
- Engineer context-aware linkers that respond to tumor-specific biochemistry
- Select optimal payload matched to tumor vulnerabilities and spatial architecture
We aren't just building better ADCs. We are engineering a new standard for oncology, one where the complexity of the tumor is no longer an obstacle, but the very roadmap to its elimination. The future of ADCs is here.
References
- Traeuble et al, Spatially Integrated Multi-Omics reveals the Multicellular Landscape of Progenitor-Driven Glioblastoma Progression. BiorXiv. February 28, 2026.
- Caroline Hoffmann, et al. “MOSAIC: Intra-tumoral heterogeneity characterization through large-scale spatial and cell-resolved multi-omics profiling” bioRxiv 2025.05.15.654189 (2025)
- Herpin et al, Deciphering Cellular Ecosystems Driving Tumor Progression and Immune Escape from Spatial Transcriptomics and Single-Cell with COMPOTES. BiorXiv. October 27, 2025.
- Fidon et al, Histology and spatial transcriptomic integration revealed infiltration zone with specific cell composition as a prognostic hotspot in glioblastoma. BiorXiv. February 12, 2026.
- Grouard et al, Multiple instance learning with spatial transcriptomics for interpretable patient-level predictions: application in glioblastoma. BiorXiv. October 15, 2025.




