Towards AI Scientists that learn from the lab
Building AI agents that don't just analyze data - they design experiments, learn from results, and iterate. A step towards closing the loop between computation and the laboratory.
TL;DR - We tested whether AI agents can learn from real experimental feedback to accelerate find which gene lies behind phenotypes induced by knock-out perturbations. Agents that received iterative results identified 166-185% more perturbation-inducing genes than random selection, demonstrating that frontier AI models can actively guide laboratory experiments, not just analyze data after the fact.
The Vision
Drug discovery is slow, expensive, and full of dead ends. Identifying the right biological targets requires vast experimental campaigns - systematically perturbing thousands of genes or compounds and measuring what happens to cells.
What if an AI system could guide this process? Not just analyze results after the fact, but actively decide which experiments to run next, learn from each round of outcomes, and progressively zero in on the most promising targets. This is the idea behind lab-in-the-loop AI: an AI scientist agent that iterates with real biological experiments, closing the loop between computation and the wet lab.
At Owkin, we set out to test whether today's frontier AI models are ready for this role. The results suggest they are closer than many expected — but also that the path forward requires careful attention to model capability and experimental design.
Why Gene-Phenotype Discovery Matters
At the heart of many diseases lies a simple question: which genes, when disrupted, change how cells behave?
When you knock out a gene and observe dramatic changes in cell shape, size, or internal structure, you have found a potential drug target. A gene whose disruption causes cancer cells to stop dividing could become the basis for a new therapy. A gene essential for bacterial survival could be the target of a new antibiotic. A gene that rescues a diseased cellular phenotype could point toward treatments for rare genetic disorders.
The challenge is scale. The human genome contains roughly 20,000 genes, and testing each one experimentally is costly. An intelligent agent that can prioritize which genes to test — and learn from each round of results to refine its next choices — could dramatically accelerate this process.
The JUMP Cell Painting Dataset
To test our AI agents, we use JUMP Cell Painting, the largest public morphological profiling resource available today.
Cell Painting is a high-content imaging assay developed at the Broad Institute. Cells are stained with six fluorescent dyes, each targeting a different organelle (nucleus, endoplasmic reticulum, mitochondria, cytoskeleton, Golgi, and plasma membrane). High-resolution microscopy captures images of thousands of cells per condition, and computational pipelines extract thousands of quantitative morphological features — measurements of cell size, shape, texture, intensity, and spatial organization.
The JUMP consortium applied this assay to ~8,000 CRISPR gene knockouts, creating a comprehensive map of how each gene perturbation alters cellular morphology. For each gene and each morphological feature, a statistical test determines whether the knockout significantly changes that feature. A "hit" means the gene knockout produces a statistically significant effect (p < 0.05) on a given cellular feature.
Our task: given a target morphological feature, discover as many gene hits as possible within a fixed experimental budget. The AI agent gets 10 rounds of 100 gene selections each (1,000 total tests from ~8,000 available genes). After each round, the agent sees which genes were hits and which were misses, and can update its strategy for the next round.
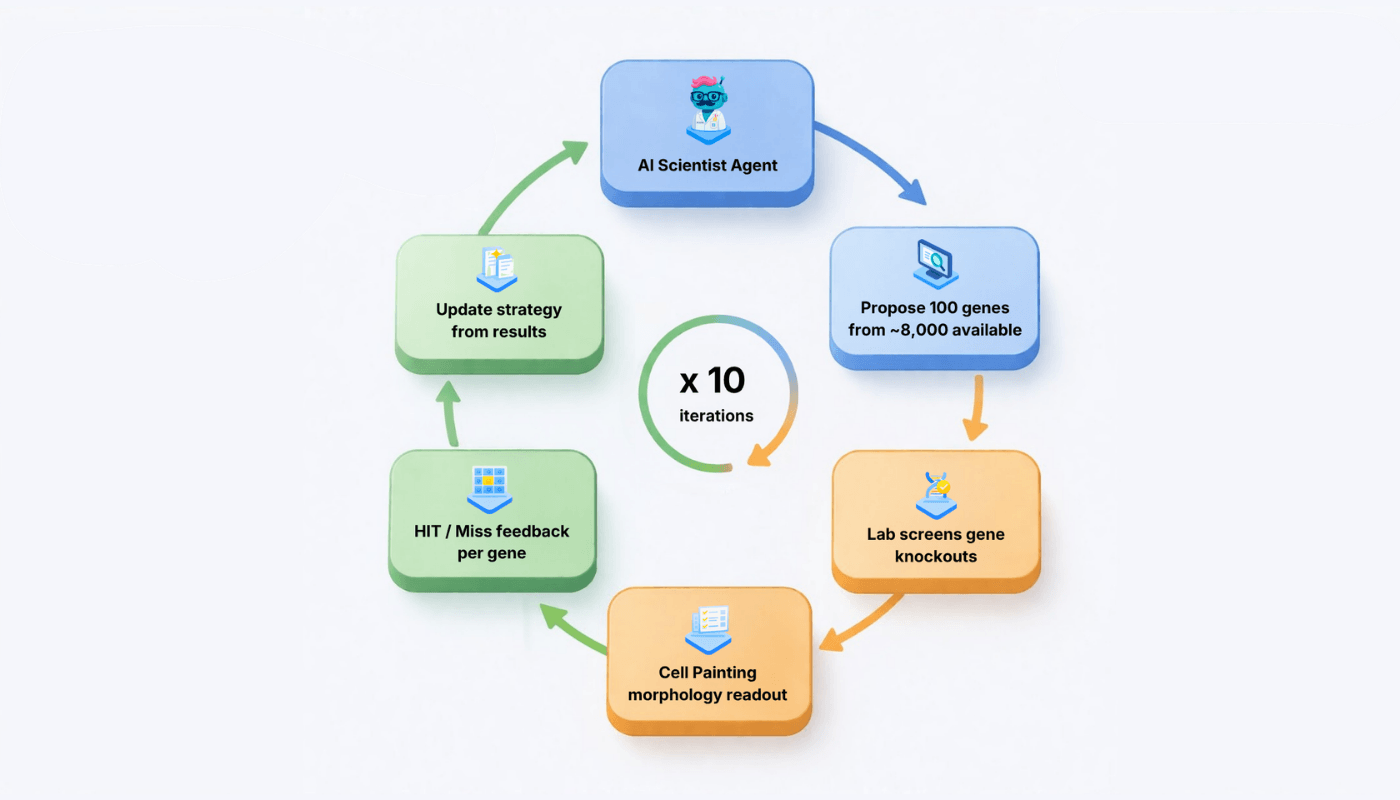
Our AI Scientist Agents
We built several agent architectures to isolate the contribution of different information sources:
- Zero-shot: The agent receives only the target feature description and the list of available genes. It selects perturbations based solely on its encoded biological knowledge — no experimental feedback whatsoever.
- ICL-EF (In-Context Learning from Experimental Feedback): Same agent, but after each round it sees the full experimental history: which genes were tested and whether they were hits or misses, along with p-values. This enables in-context learning (ICL) — the agent can reason about patterns in the results.
- ICBR-EF (In-Context Belief Revision from Experimental Feedback): Extends ICL-EF by also receiving quantitative morphological signatures (the top 10 most affected features) for a selection of tested genes (8 most recent hits and 4 most recent misses) and maintaining a structured hypothesis register that tracks the lifecycle of biological theories across iterations.
We also compare against two non-LLM baselines:
- Random: genes chosen uniformly at random.
- GP-UCB: a classical Gaussian Process optimizer using protein-protein interaction networks from STRING as its similarity kernel — a similarity function that tells the algorithm how related two genes are. GP-UCB is a standard Bayesian optimization approach that also receives iterative feedback.
All LLM agents use Claude Sonnet (versions 4.5 or 4.6). We run 10 replicates for each of 10 target features across 8 conditions — 800 total experiments.
Results deep dive: The Agent Learns from Feedback
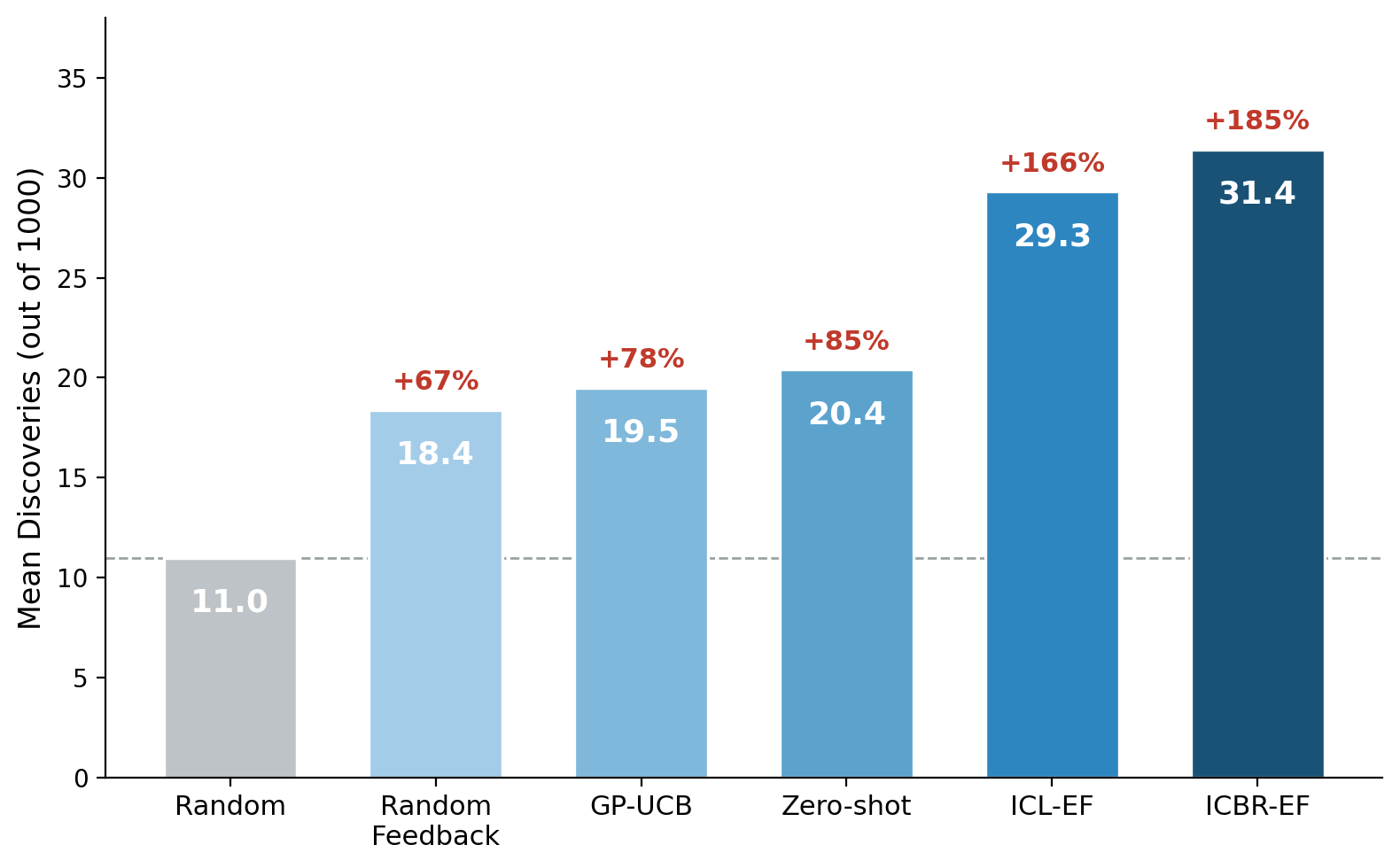
The zero-shot agent — using only its biological priors — already beats random selection by 85%, confirming that frontier models encode substantial and useful biological knowledge.
But the ICL-EF agent, which receives experimental feedback, performs substantially better: 29.3 mean discoveries (+166% over random), gaining +8.9 discoveries per feature over the no-feedback baseline (p = 0.003). The ICBR-EF agent, which additionally receives phenotypic signatures, achieves 31.4 — and the marginal gain over ICL-EF (+2.1) is statistically significant (p = 0.006).
Both feedback-enabled agents outperform GP-UCB (19.5, p = 0.003), suggesting that LLM-based semantic reasoning can be more effective than statistical regression for this task. We note, however, that GP-UCB used a STRING PPI kernel, which measures similarity based on protein interactions rather than morphological effects. Since genes from different pathways can produce similar morphologies, and vice versa, this may not be the optimal similarity measure for this task. Richer kernels (e.g., based on gene expression patterns or morphological embeddings) could narrow this gap.
The Random Feedback Control
To confirm the agent genuinely learns from the content of feedback (not just from having a longer prompt), we ran a Random Feedback control: the agent receives feedback where hit/miss labels are randomly permuted, destroying the information content while preserving feedback structure.
Random feedback doesn't help — it hurts (-2.0 vs no feedback, harmful on 6/10 features). The agent chases phantom signals, wasting its budget on misleading leads. The full ICL effect (+13.0 for ICBR-EF over permuted feedback, p = 0.003) is attributable to genuine learning from the factual content of results.
How the Agent Learns
Analysis of the agents' reasoning traces reveals a clear evolution in strategy:
From static priors to pathway exploitation. The zero-shot agent makes the same gene selections regardless of outcomes. The ICL-EF agent, by contrast, pivots once it observes hits: if it discovers a hit in the SWI/SNF chromatin remodeling complex, it immediately targets the remaining untested members of that complex. Semantic analysis shows it references specific "complex/family/subunit" exploitation 7.9 times per iteration, versus 2.3 for the zero-shot agent (a 3.4× higher rate).
From pathway exploitation to phenotypic state inference. The ICBR-EF agent goes further. Because it receives multidimensional morphological fingerprints, it can cluster hits by their cellular effects rather than their gene names. For example, on feature F80 (cytoplasm granularity), the agent observed that MDM2 (a p53 regulator), WEE1 (a G2/M checkpoint kinase), and TTK (a spindle assembly checkpoint kinase) — functionally distinct genes — all shared the same side-effect signature in AGP texture features (AngularSecondMoment, DifferenceEntropy, InverseDifferenceMoment). It deduced they converged on a common physiological state — mitotic arrest causing cytoplasmic reorganization — and targeted additional cell cycle regulators from that phenotypic cluster. It references phenotypic states 22.6 times per iteration (vs. 0.7 for ICL-EF).
Model Capability Matters
We ran the same experiments with Claude Sonnet 4.5 (200 additional replicates). The contrast is striking:
With Sonnet 4.5, nearly half of proposed genes are real biological genes that don't exist in the screening library — for instance, EXOSC1 (a real exosome subunit) when the library only contains EXOSC4-9, or MED18 (a real Mediator subunit) absent from the library. This is not random noise — it is an instruction-following failure: the model proposes biologically sensible genes from its training knowledge rather than constraining its output to the provided list. The problem worsens over iterations (from 0% at iteration 1 to ~60% by iteration 6, plateauing thereafter) as the untested list shrinks and the model increasingly defaults to its priors.
Sonnet 4.6 reliably constrains its output to the provided gene list, reducing out-of-library proposals ~5×. This allows the agent to use its full budget for intentional selections, and discoveries jump from 21.8 to 29.3 (+34%).
An important caveat: a model upgrade changes many capabilities simultaneously — world knowledge, reasoning, instruction adherence — so the improvement cannot be attributed to any single factor. What we can conclude is that sufficient model capability is necessary for ICL to manifest.
The Road Ahead: From Retrospective Data to Real Experiments
Our results are encouraging, but they come with an important caveat: all experiments use pre-computed p-values from the JUMP dataset. The feedback is clean, binary, noise-free, and instantaneous. Real experimental campaigns involve batch effects, measurement noise, variable hit rates across plates, and days or weeks of latency between proposing an experiment and seeing results.
The real test of AI scientist agents will come when they are deployed in actual lab-in-the-loop settings — proposing experiments that are physically executed, receiving noisy real-world feedback, and adapting under genuine uncertainty. Whether the ICL advantage we observe survives the transition from retrospective simulation to live experimentation remains an open question — and the most important one.
Beyond in-context learning, reinforcement learning offers a natural extension: training the agent directly on experimental outcomes to develop persistent, optimizable policies. The JUMP dataset provides a simulated environment for this training, and the behavioral patterns we observe (aggressive pathway exploitation vs. unproductive rabbit holes) map directly onto the exploration-exploitation trade-off that RL is designed to optimize.
Cost
Each 10-iteration campaign requires 10 LLM API calls, consuming ~410K input tokens and ~19K output tokens. At current Claude Sonnet pricing ($3/MTok input, $15/MTok output), that's roughly $1.50 per campaign ($1.41 for zero-shot, $1.46 for ICL-EF, $1.77 for ICBR-EF), or about $0.05 per discovery. The full 600 LLM campaigns in this study cost approximately $900 in API calls.
Caveats
Several important caveats apply. All experiments use pre-computed data; real experiments would introduce noise that could alter ICL dynamics. We evaluate only two proprietary Claude models — generalization to other models is unknown. The Sonnet 4.5 to 4.6 comparison is confounded: the upgrade changes world knowledge, reasoning quality, instruction adherence, and hallucination simultaneously, so the ICL improvement cannot be attributed to any single factor. Frontier LLMs may have seen JUMP-related papers during training, potentially inflating the prior-knowledge baseline. And we do not evaluate the LLMNN hybrid method proposed by Gupta et al., which delegates iterative updates to a classical nearest-neighbor algorithm.
What This Means for Owkin
At Owkin, we are building AI systems that work alongside scientists to accelerate drug discovery. K Pro, our AI scientist, is evolving to integrate agentic capabilities that can analyze complex datasets and generate hypotheses about disease mechanisms and therapeutic targets. But generating hypotheses is only half the problem; validating them experimentally is the bottleneck.
This work demonstrates that frontier AI agents can do more than retrieve biological knowledge — they can learn from experimental outcomes and progressively refine their strategies when model capability is sufficient. When AI scientists like K Pro are plugged into automated or semi-automated laboratory systems, this capability will enable them to select experiments, read results, interpret outcomes, and iteratively propose the next round of perturbations. The vision is a closed loop between computational hypothesis generation and experimental validation.
The stakes are high. An AI scientist that can efficiently guide perturbation screens — identifying cancer vulnerabilities, antibiotic targets, or disease mechanisms — could substantially accelerate the path from biological hypothesis to therapeutic candidate. Our results, while specific to one dataset and two models, suggest that the foundations for lab-in-the-loop AI are being laid today. The next step is testing these capabilities in live experimental settings, where they will face the noise and complexity of real biology.
Full results are available as a pre-print on arxiv at: https://arxiv.org/abs/2603.26177
This work was conducted across 800 independently replicated experiments (8 agent configurations × 10 features × 10 replicates, including both Claude Sonnet 4.5 (claude-sonnet-4-5-20250929) and 4.6 (claude-sonnet-4-6)). The code and benchmarks are available at github.com/owkin/frontier-wp5.




