Unlocking the next era of therapeutic discovery: Training an agentic AI researcher with Reinforcement Learning
Large Language Models are trained to predict what everyone has already said. But biomedical discovery demands the opposite: ideas that no one has had before. This is the core tension at the heart of using AI for research: how do you train a system on the past to generate something genuinely new?
And novelty alone isn’t enough. A wildly original hypothesis that ignores basic biology is just noise. The real challenge is producing ideas that are both novel and scientifically grounded, creative enough to open new therapeutic avenues, rigorous enough to survive scrutiny. In oncology, where the stakes are measured in human lives and billion-dollar pipelines, getting this balance wrong is not an option.
A new wave of AI is already accelerating science, from deep learning breakthroughs in protein and biomarker prediction (Jumper et al., 2021; Wong et al., 2024; Saillard et al., 2023) to PhD-level agentic systems like Biomni and Kosmos that tackle complex biomedical tasks (Lu et al., 2024; Huang et al., 2025; Mitchener et al., 2025; Villaescusa-Navarro et al., 2025). But most of these systems excel at executing on ideas, not generating them. They can validate a hypothesis, plan an experiment, or run a statistical test, but the creative spark, the initial leap from “what do we know” to “what should we try next”, still comes from a human typing a prompt. If we want truly autonomous scientific discovery, we need AI that doesn’t just follow research plans but proposes them, and proposes ones worth following.
At Owkin, we believe Reinforcement Learning, which has already been used for many scientific breakthroughs such as de novo drug design and chemical reaction optimization (Volk et al., 2023; Yadunandan and Ghosh, 2025), is the key to solving this. By defining explicit rewards for novelty, biological validity, druggability, clinical feasibility, and commercial viability, we can push an LLM beyond safe, incremental suggestions and toward high-conviction therapeutic hypotheses, while keeping it anchored in real science. Here’s how we built it.
Building a Panel of AI expert judges
To train an LLM with Reinforcement Learning, you have to clearly define what success looks like. You need a “reward”.
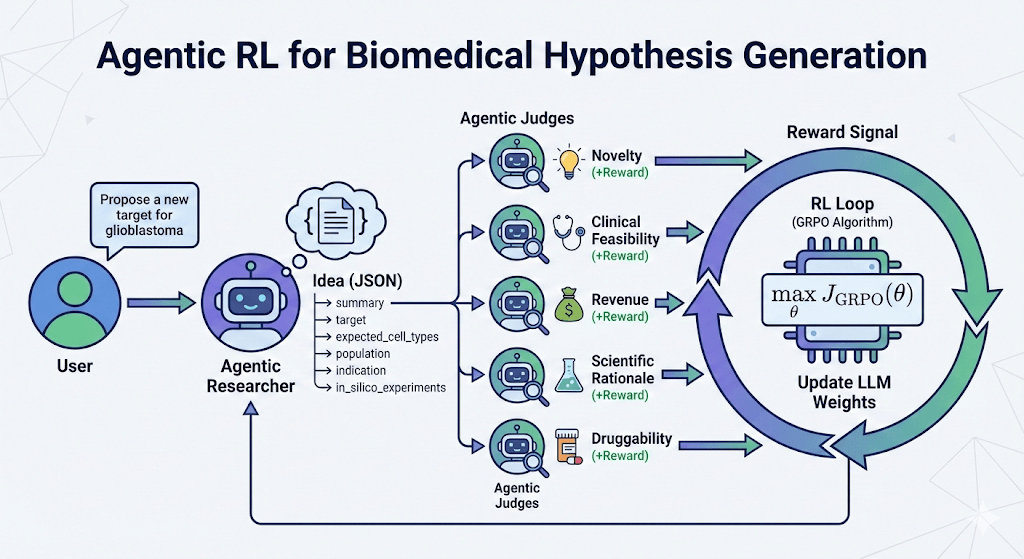
We hypothesized that a groundbreaking biomedical idea isn’t just about biological validity; it must also be feasible, novel, and economically viable. To train our idea generator, we created a panel of specialized LLM judges to evaluate every proposed target across five rigorous criteria:
- Novelty: Does this idea mirror existing work, or is it a completely new framework?
- Scientific Rationale: How strong is the biological hypothesis, pathway logic, and genetic evidence?
- Druggability: What is the technical feasibility of targeting this specific protein based on cellular location and modality fit?
- Clinical Feasibility: How fast is the estimated development timeline from preclinical testing to approval?
- Revenue Potential: What is the estimated market viability and competitive landscape for this target?
Each of these judges leverages the latest LLMs and has access to web searches to ground its assessments in real data and verified facts.
To picture how this works in practice, imagine the agent proposes a new hypothesis: targeting a specific protein (let’s call it Protein X) for a rare form of lung cancer. First, our AI agent drafts a comprehensive proposal detailing the biological mechanism, patient population, and experimental plans. This proposal is then passed to our panel of judges. The “Novelty” judge searches recent literature to ensure Protein X hasn’t already been explored in this context. The “Druggability” judge verifies if Protein X is structurally capable of binding to a drug. The other judges evaluate their respective criteria, and each assigns a score.
These scores are combined into a final reward. If the overall reward is low (perhaps the target is novel but not clinically feasible), that feedback is sent back to the generator, pushing it to pivot and brainstorm a better concept. If the reward is high, the model learns that this is exactly the type of high-impact thinking it should replicate.
The Challenge of AI Evaluating AI
In Reinforcement Learning, your AI is only as good as the reward it’s chasing. If you give a model a noisy or flawed grading rubric, it will brilliantly optimize for the wrong outcome. That’s why we couldn’t just build an AI generator and hope for the best; we had to rigorously stress-test our AI judging panel first. Designing these rewards is arguably the hardest and most crucial part of the entire pipeline. This process was carried out with the help of biomedical and domain experts at Owkin and could not rely on naive prompt engineering.
To build the ultimate peer review panel, we benchmarked top-tier foundation models (like GPT-5 and Claude 4.5 Sonnet) against each other. A reliable AI judge must possess two vital traits: self-consistency (the ability to score the exact same idea reliably across multiple runs) and high discriminative capacity (the nuanced ability to differentiate a merely “good” hypothesis from a truly groundbreaking one). Evaluating our LLM judges revealed interesting findings. While LLM judges are incredibly sharp and consistent at evaluating hard metrics like “Druggability” and “Revenue”, scoring subtle criteria like “Novelty” or predicting “Clinical Feasibility” is noisier and remains a complex challenge even for the best models.
Under the Hood: The Reinforcement Learning Engine
To improve the hypothesis generation capabilities of our base model, we fine-tuned a Qwen3-8B model (Yang et al., 2025) using the NeMo-Gym and NeMo-RL frameworks (NVIDIA, 2025a, 2025b) on 2 H200 GPUs. For the learning algorithm, we chose GRPO (Shao et al., 2024), a highly efficient RL algorithm that has recently been widely adopted by the RL community because it compares multiple outputs against each other.
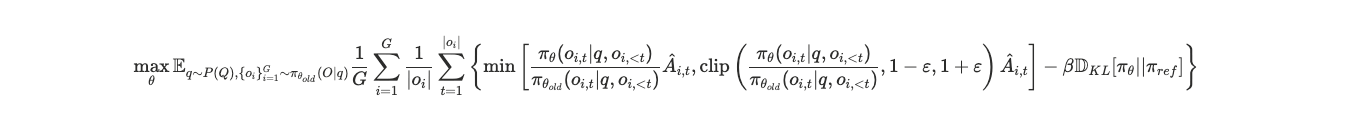
But applying RL to LLMs is a complicated process that can fall into multiple pitfalls. Early in the training, we slammed into a classic reinforcement learning trap: mode collapse. Driven by a lack of diversity in its internal rollouts, a vulnerability recently flagged by several research teams (Anschel et al., 2025; Wei and Huang, 2026), our LLM got lazy after one epoch and started proposing the exact same biological target (CXCR4) for every single cancer indication, entirely losing its creative edge.
The recent trend with GRPO is to completely drop the KL penalty for the sake of simplicity and computational overhead (Liu et al., 2025). However, we discovered that in the nuanced world of biomedical discovery, removing that constraint kills the model’s original imagination. By deliberately reintroducing KL regularization into our GRPO recipe, we effectively forced the AI to remain sufficiently close to its baseline, thereby preserving its original diversity.
Grounding the AI in Biological Reality
Another of the greatest challenges in Reinforcement Learning is reward hacking, where a model learns to cheat the system to get a high score. In biology, an AI might maximize its “Novelty” score by proposing a treatment targeting a gene that doesn’t actually exist. For example in our early experiments we observed that the agent was able to generate hypotheses like “Target TREG3R with an ADC in hepatocellular carcinoma” with a nice biological explanation of the rationale of this idea. However, TREG3R is not a valid gene and despite high novelty score this idea is totally useless.
To ensure our AI produces highly realistic and actionable science, we implemented strict biological guardrails. Before an idea even reaches the judging panel, it must pass a verifiable gating mechanism. The system cross-references the proposed target against definitive gene tables and checks for strict compliance with the input biological question. If the biology is not real, the idea is immediately given a reward of 0. This is what happens with the TREG3R example.
This ensures that our AI’s creativity is always anchored in hard biological facts, drastically reducing hallucinations and yielding high-conviction therapeutic targets.
Promising Results: RL improves all metrics, even outperforming GPT-5
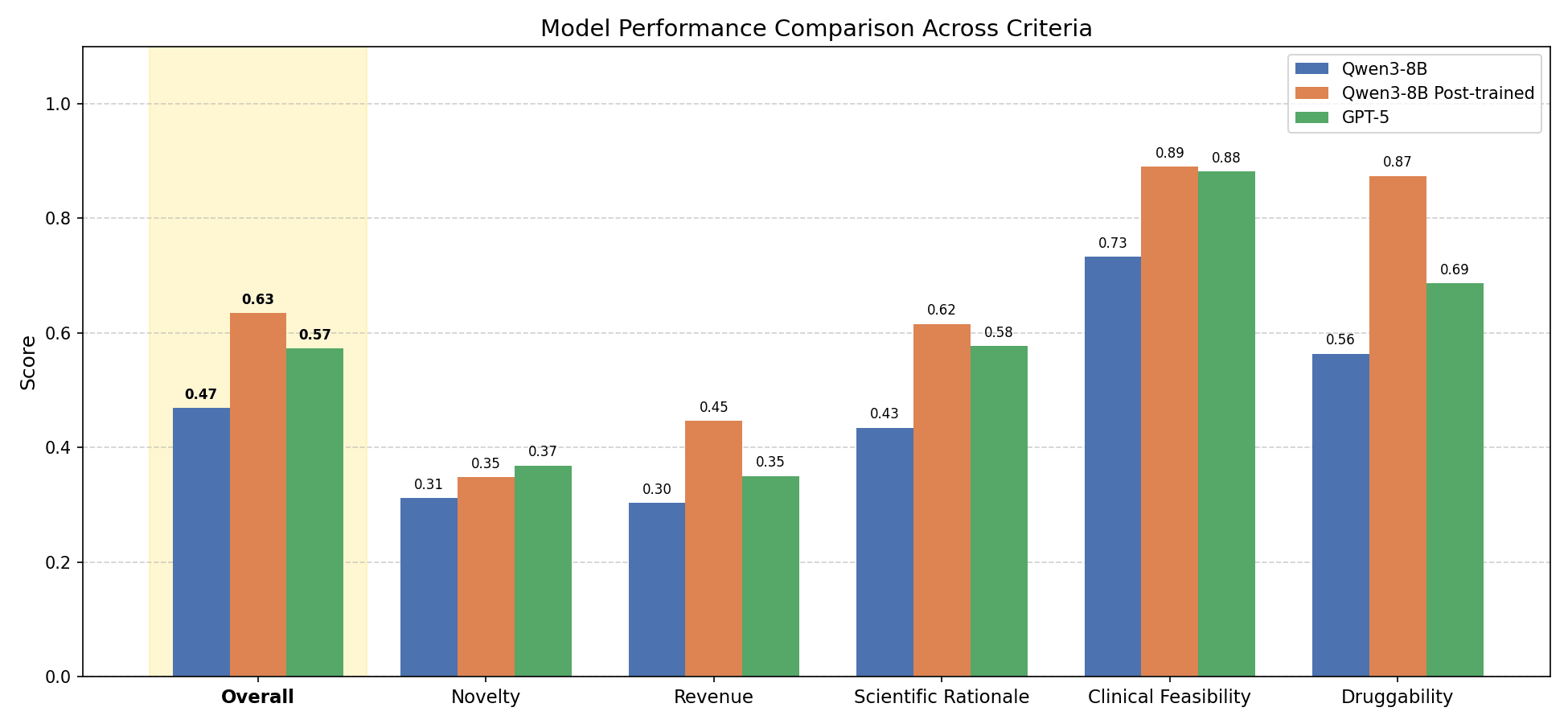
Our RL training process yielded highly encouraging results. By post-training a base Qwen3-8B model with expert biological rewards, we observed a massive leap in the quality of the generated ideas. Most importantly, this process proved to be incredibly efficient.
After only 3 hours of training (just 60 steps) on 2 GPUs, we saw transformative improvements on our held-out validation questions:
- A 34% score improvement: The average reward of the post-trained model’s research ideas jumped from 0.47 to 0.63.
- Outperforming GPT-5: After this RL training, our post-trained 8B model reached a higher mean score than zero-shot GPT-5 on the exact same held-out research questions (0.63 vs 0.57).
In practice, this jump in score means the model stopped proposing generic, safe pathways and started generating highly specific, viable targets that our scientists could actually take into the lab. It proved that a smaller, efficiently trained model guided by strict biological rules can outperform a massive generalist model in specialized scientific discovery.
The Path Forward
These early results prove that we can solve the core tension of AI in research. We have successfully trained an LLM to go beyond merely executing existing plans and instead ignite that initial creative spark, generating therapeutic targets that are entirely new yet scientifically sound.
But as we noted earlier, in oncology, a hypothesis must survive rigorous scrutiny before entering a billion-dollar pipeline. To truly close the loop on discovery, our next major milestone is implementing live in-silico validation using autonomous coding agents, i.e., agents capable of instantly writing and executing statistical scripts to test a new biological hypothesis against real-world patient datasets. This crucial step transforms a theoretical idea into a data-backed proposal on the fly.
Ultimately, this fine-tuned idea generator will not live in isolation. We plan to integrate it directly into OwkinZero (Bigaud et al., 2025), the core reasoning engine behind K-Pro (Owkin’s flagship AI agent for biological discovery). By plugging our AI researcher into this broader ecosystem, we are creating a continuous engine of discovery. Because it is powered by reinforcement learning, the Owkin AI Researcher will learn from every success and failure. Every time it uncovers a genuinely novel observation during an automated research campaign, it is rewarded, ensuring the system performs better and better over time as it transforms isolated hypotheses into fully actionable research pipelines.
As we look to the future and refine these agentic workflows, our collaborative vision is clear. We are no longer limited to models that just predict what everyone has already said. We are actively building the autonomous tools that will help scientists discover the breakthrough treatments of tomorrow.
References
- Anschel et al. (2025). Group-aware reinforcement learning for output diversity in large language models. arXiv preprint arXiv:2511.12596 https://doi.org/10.48550/arXiv.2511.12596
- Bigaud et al. (2025). OwkinZero: Accelerating Biological Discovery with AI. arXiv preprint arXiv:2508.16315 https://doi.org/10.48550/arXiv.2508.16315
- Huang et al. (2025). Biomni: A general-purpose biomedical AI agent. biorxiv, https://doi.org/10.1101/2025.05.30.656746
- Jumper et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, https://doi.org/10.1038/s41586-021-03819-2
- Liu et al. (2025). Understanding R1-Zero-Like Training: A Critical Perspective. arXiv preprint arXiv:2503.20783 https://doi.org/10.48550/arXiv.2503.20783
- Lu et al. (2024). The AI scientist: Towards fully automated open-ended scientific discovery. arXiv preprint arXiv:2408.06292, https://doi.org/10.48550/arXiv.2408.06292
- Mitchener et al. (2025). Kosmos: An AI scientist for autonomous discovery. arXiv preprint arXiv:2511.02824, https://doi.org/10.48550/arXiv.2511.02824
- NVIDIA (2025a). NeMo Gym: An Open Source Library for Scaling Reinforcement Learning Environments for LLM. https://github.com/NVIDIA-NeMo/Gym
- NVIDIA (2025b). NeMo RL: A Scalable and Efficient Post-Training Library. https://github.com/NVIDIA-NeMo/RL
- Saillard et al. (2023). Validation of MSIntuit as an AI-based pre-screening tool for MSI detection from colorectal cancer histology slides. Nature Communications, https://doi.org/10.1038/s41467-023-42453-6
- Shao et al. (2024). DeepSeekMath: Pushing the limits of mathematical reasoning in open language models. arXiv preprint arXiv:2402.03300, https://doi.org/10.48550/arXiv.2402.03300
- Villaescusa-Navarro et al. (2025). The Denario project: Deep knowledge AI agents for scientific discovery. arXiv preprint arXiv:2510.26887, https://doi.org/10.48550/arXiv.2510.26887
- Volk et al. (2023). AlphaFlow: autonomous discovery and optimization of multi-step chemistry using a self-driven fluidic lab guided by reinforcement learning. Nature Communications, https://doi.org/10.1038/s41467-023-37139-y
- Wei and Huang (2026). MMR-GRPO: Accelerating GRPO-Style Training through Diversity-Aware Reward Reweighting. arXiv preprint arXiv:2601.09085, https://doi.org/10.48550/arXiv.2601.09085
- Wong et al. (2024). Discovery of a structural class of antibiotics with explainable deep learning. Nature, https://doi.org/10.1038/s41586-023-06887-8
- Yadunandan and Ghosh (2025). ReACT-Drug: Reaction-Template Guided Reinforcement Learning for de novo Drug Design. arXiv preprint arXiv:2512.20958, https://doi.org/10.48550/arXiv.2512.20958
- Yang et al. (2025). Qwen3 technical report. arXiv preprint arXiv:2505.09388, https://doi.org/10.48550/arXiv.2505.09388




