Teaching AI with reinforcement learning in a “biology video game” to find the root cause of disease
Can LLMs Think Like Biologists? Teaching AI to Reverse-Engineer Gene Networks
When a gene expression shows dramatic changes in disease, is it the root cause—or just a bystander caught in a cascade? Distinguishing causes from consequences remains one of drug discovery's most expensive unsolved problems. We trained a mid-sized open-source AI model using reinforcement learning on 20,000 synthetic gene network puzzles. The question: could we teach it to reason like a biologist—tracing effects upstream to find the hidden driver?

From Symptoms to Sources: The Quest for Root Causes in Medicine
The ultimate ambition of biomedical research is to find the root causes of disease and develop treatments that cure the disease itself — not merely its symptoms. For over a century, physicians treated peptic ulcers with antacids and stress management, until Marshall and Warren discovered in 1982 that the true culprit was a bacterium, Helicobacter pylori, transforming a chronic condition into one curable with antibiotics (Marshall & Warren, 1984; Nobel Prize, 2005). What the medical community had long treated as a consequence — excess acid, inflammation — was masking a far simpler cause. This confusion between cause and consequence haunts biomedicine to this day. Consider Alzheimer's disease: for over thirty years, the amyloid cascade hypothesis has driven billions in research spending and dozens of clinical trials, yet whether amyloid-β plaques are a root cause or a downstream bystander remains fiercely debated (Hardy & Higgins, 1992; Herrup, 2015; Behl, 2024).
Drowning in Data: The Curse of Correlation in Modern Transcriptomics
The revolution in next-generation sequencing and single-cell transcriptomics has given researchers extraordinary tools to tackle this challenge, enabling the measurement of tens of thousands of genes in individual cells (Tang et al., 2009). This has yielded spectacular successes — the discovery that chronic myeloid leukemia is driven by the BCR-ABL fusion gene led directly to imatinib, transforming a fatal cancer into a manageable condition (Druker et al., 2001). Yet for every imatinib, countless targets that looked promising in expression analyses crumbled in clinical trials because they were consequences, not causes. When you measure thousands of variables simultaneously, disentangling cause from correlation becomes exponentially harder.
Disentangling the Avalanche: Tracing Downstream Signals Back to Their Source
To make this concrete, imagine you compare gene expression in tumor versus healthy tissue and observe that hundreds of genes differ dramatically. Which one is the root cause? Your first instinct — pick the gene that changes the most — is reasonable but often wrong. In biological networks, a subtle perturbation of one upstream "master regulator" can trigger an avalanche of downstream changes, and the loudest signals are often the farthest downstream effects. If you could trace the network of regulatory influences back upstream, you might find the single perturbation that explains everything. But real gene regulatory networks involve thousands of interconnected genes with dense feedback loops, making this abductive reasoning problem — inferring the best explanation from observations — far beyond simple intuition.
Enter the Reasoning Gym: A Biological "Video Game" to Train AI
What if AI could help? At Owkin, we believe this is one of the central challenges for the application of modern AI to biomedical research, and that progress here can have immediate impact for millions of patients. Motivated by the recent success of reinforcement learning in training large language models for advanced reasoning — from OpenAI's o1 (OpenAI, 2024) to DeepSeek-R1 (Guo et al., 2025), which showed that LLMs can develop sophisticated reasoning purely through RL — we designed an original biological reasoning simulation environment. In AI parlance, a reasoning gym: a video game of the cell, played by AI, where the model faces thousands of "root cause finding" challenges with known ground truth, and is rewarded when it reasons correctly.
Constructing Causal Puzzles: Benchmarking AI on Gene Regulatory Networks
To operationalize this environment, we focus on a specific challenge: given a known gene regulatory network (GRN) and the expression state before and after a hidden intervention, can an LLM infer which single regulatory relationship was perturbed?
Real-world GRNs are notoriously difficult to reconstruct (Mao et al., 2022). To isolate pure reasoning ability from experimental noise, we use synthetically generated GRNs, a standard benchmarking strategy (Saint-Antoine & Singh, 2023; Pratapa et al., 2020).
Our simulator generates ground-truth causal puzzles at scale through three steps:
- Simulation: We generate a GRN and simulate its dynamics until it reaches a stable baseline state.
- Intervention: We perturb a single connection (edge) in the network and simulate the system again until it reaches a new "perturbed" state.
- Inference: The model is shown the network topology and the change in gene expression between the two states. Crucially, the perturbation itself is hidden. The model must deduce which specific edge was modified to produce the observed shift.
By controlling the parameters of the simulation, we can generate thousands of these "causal problems" to train and test the model's reasoning capabilities.
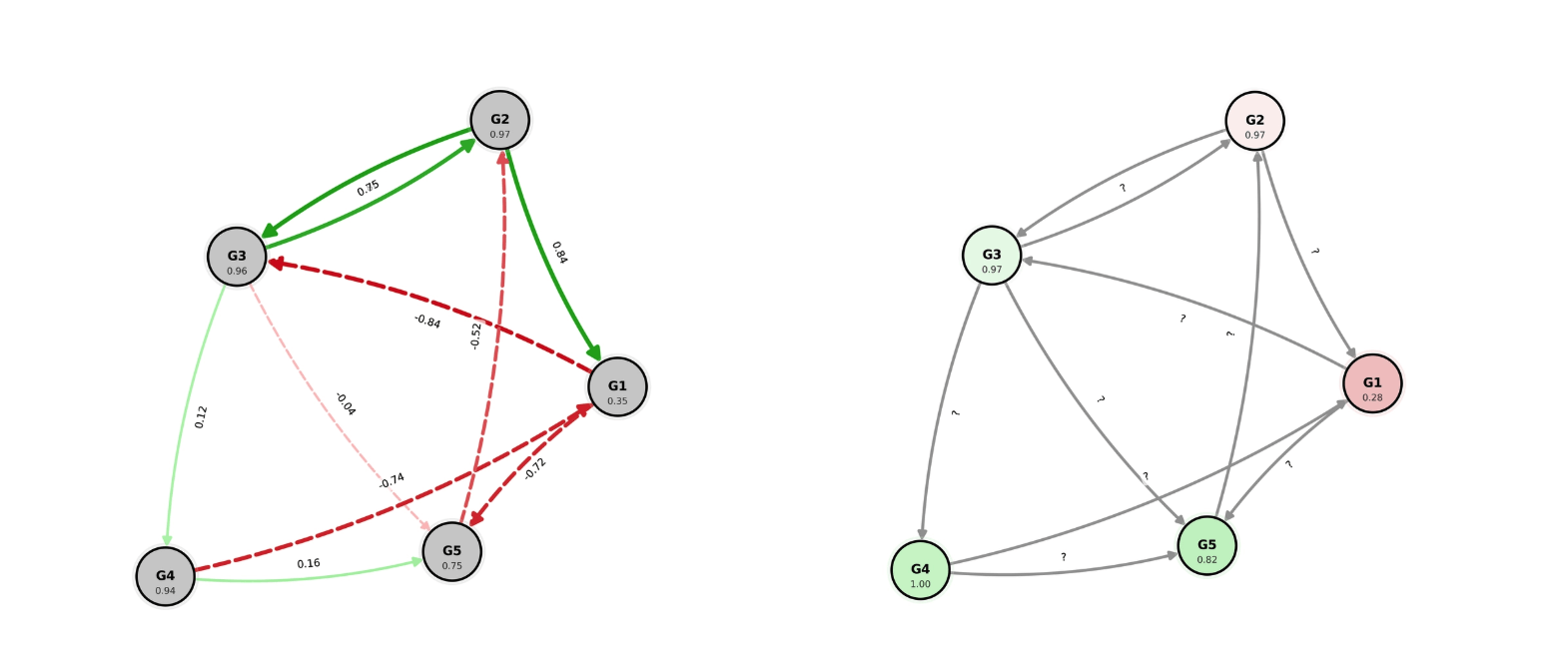
Right: Graph after perturbation as seen by LLMs: the expression change is seen but the new edge weights are hidden for LLMs to retrieve the cause of the perturbation.
Designing the Challenge: Distinguishing Local Effects from Downstream Avalanches
To recreate the "avalanche" effect described earlier — where the true root cause is masked by loud downstream signals — we tuned the steepness of our simulator's gene activation functions (Vohradský, 2001). This creates a regime where indirect effects dominate the expression changes.
We generated 20,000 puzzles split into two difficulty tiers:
- Local effects: The target gene changes the most (solvable by heuristics).
- Propagated effects: The target is masked by downstream avalanches.
Propagated effects are the true test: to solve them, the model cannot just "pick the biggest number"—it must trace the causal chain upstream through feedback loops and regulation.
Post-Training Open LLMs: Using Reinforcement Learning for Biological Inference
We post-trained open-weight LLMs (Qwen3 family) using reinforcement learning (RL) on 8 NVIDIA H200 GPUs. The reward function explicitly incentivizes identifying the hidden perturbed edge, with higher rewards for correctly pinpointing the direct target gene.
We compared Group Relative Policy Optimization (GRPO) with Group Sequence Policy Optimization (GSPO) (Zheng et al., 2025). While GRPO struggled with stability on our hardest graphs—sometimes entering infinite reasoning loops—GSPO proved robust, allowing us to train longer and achieve higher performance.
Results: A Mid-Sized Open Model Matches Proprietary AI in Causal Reasoning
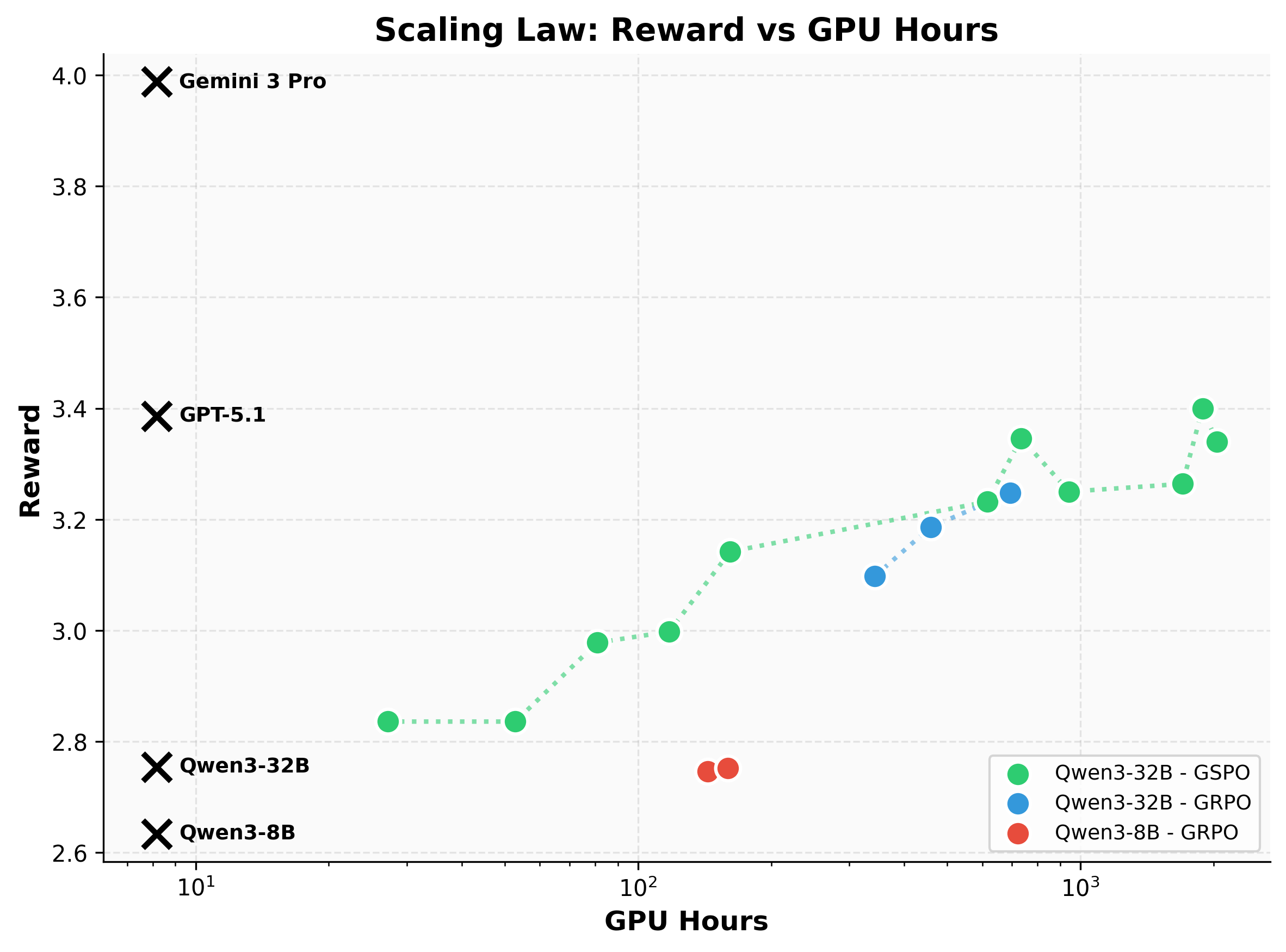
A mid-sized open model matches GPT-5.1. Our best model, Qwen3-32B post-trained with GSPO, reaches a reward of 3.4, effectively matching GPT-5.1's zero-shot performance. While Gemini achieves higher scores (~4.0), it does so as a significantly larger frontier model. This demonstrates that specialized post-training can close the gap between mid-sized open models and proprietary frontier systems on complex reasoning tasks, making this capability more accessible and deployable.
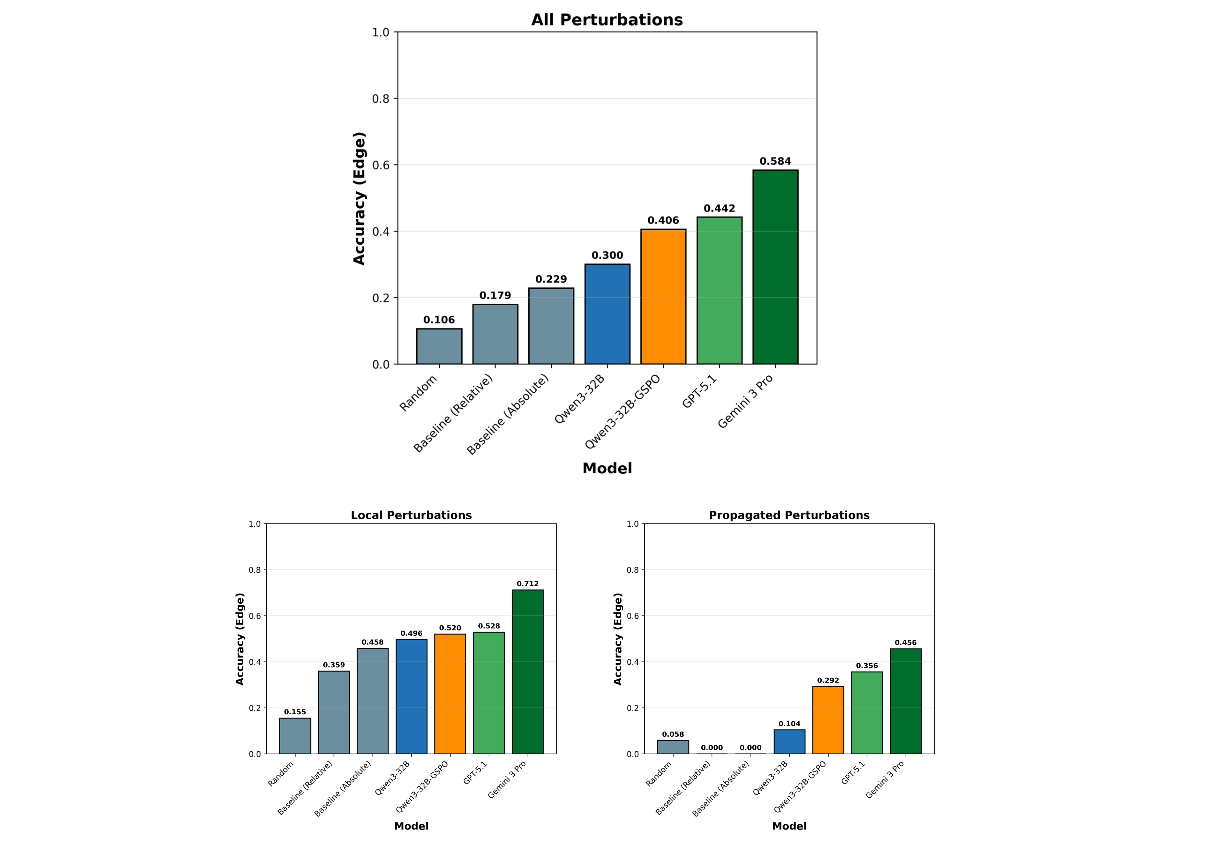
It learns to trace cascades. Base Qwen3-32B fails almost completely on propagated effects (~10% accuracy). After training, accuracy triples to 29%, while accuracy on easier local effects holds steady. The model effectively learns to ignore the "loudest" signal and hunt for the "silent" driver upstream.
For example, consider the scenario shown in the first figure. In this scenario, a perturbation was applied to the connection G3→G4, triggering a cascade, causing a downstream gene (G1) to exhibit the largest expression change (-20.5%). The base model typically falls for the decoy, incorrectly identifying the "loudest" gene (G1) as the direct target. In contrast, our post-trained model correctly identifies the true root cause (G3→G4), reasoning that the stable upstream regulator (G3) driving the intermediate changes offers the only consistent explanation for the observed propagation pattern.
Looking Ahead: Scaling from Synthetic Graphs to Real-World Disease with OwkinZero
This work validates a crucial principle: LLMs can acquire structured causal reasoning through reinforcement learning.
While much of frontier AI research focuses on general reasoning capabilities, we're proving those advances can be adapted for biology's hardest problems. This research will be integrated into OwkinZero (Bigaud et Al., 2025), the reasoning engine powering K Pro, Owkin's agentic AI platform for drug discovery. While we validated the approach on synthetic networks, the same principles apply to real patient data: identifying which molecular perturbations drive disease, not just correlate with it. As we scale these capabilities to genome-wide networks derived from patient cohorts, we're building AI that doesn't just analyze biology, it reasons about causality. The next breakthroughs in medicine may not come from generating more data, but from finally understanding the data we have.
Learn more about Owkin Zero and K Pro
References
- Marshall & Warren (1984). The Lancet, 323(8390), 1311–1315. https://doi.org/10.1016/S0140-6736(84)91816-6
- Hardy & Higgins (1992). Science, 256(5054), 184–185. https://doi.org/10.1126/science.1566067
- Herrup (2015). Nature Neuroscience, 18(6), 794–799. https://doi.org/10.1038/nn.4017
- Behl (2024). Frontiers in Aging Neuroscience, 16, 1459224. https://doi.org/10.3389/fnagi.2024.1459224
- Tang et al. (2009). Nature Methods, 6(5), 377–382. https://doi.org/10.1038/nmeth.1315
- Druker et al. (2001). NEJM, 344(14), 1031–1037. https://doi.org/10.1056/nejm200104053441401
- OpenAI (2024). o1 System Card (arXiv:2412.16720). https://doi.org/10.48550/arXiv.2412.16720
- Guo et al. (2025). Nature, 645, 633–638. https://doi.org/10.1038/s41586-025-09422-z
- Mao et al. (2022). BMC bioinformatics, 23(1), 503. https://doi.org/10.1186/s12859-022-05055-5
- Saint-Antoine & Singh (2023). BioRxiv. https://www.biorxiv.org/content/10.1101/2023.05.12.540581v1
- Pratapa et al. (2020). Nature methods, 17(2), 147-154. https://doi.org/10.1038/s41592-019-0690-6
- Vohradský (2001). the FASEB journal, 15(3), 846-854. https://doi.org/10.1096/fj.00-0361com
- Zheng et al. (2025). arXiv preprint https://doi.org/10.48550/arXiv.2507.18071
- Bigaud et al. (2025) arXiv preprint https://doi.org/10.48550/arXiv.2508.16315




